Three of the surviving subclades of mtDNA haplogroup C1 (C1b, C1c, and C1d) are found almost exclusively in the New World, with oldest sample I11974, dating back to nearly 12000 ybp (years before present) from Coquimbo, Chile.
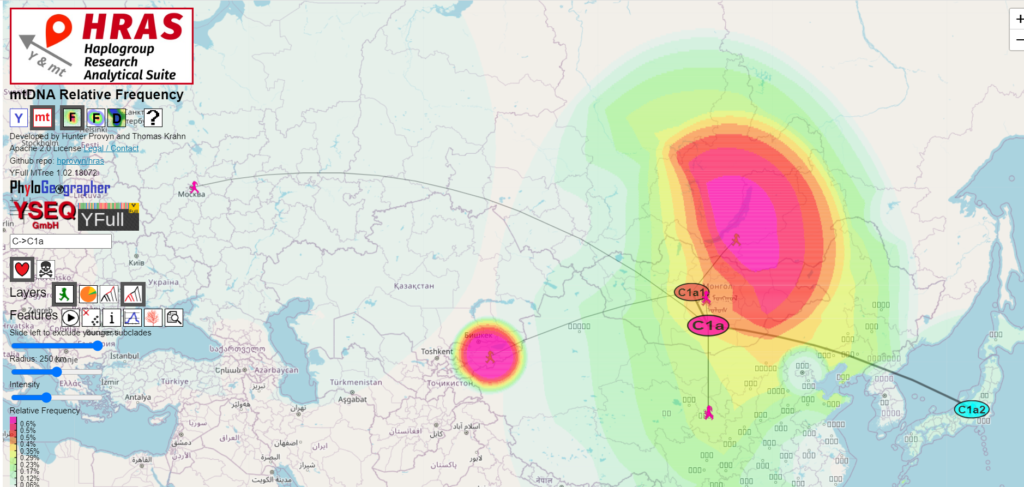
However, C1a is found exclusively in North East Asia, descending from a MRCA (Most Recent Common Ancestor) estimated by the YFull MTree to have lived about 8100 ybp (years before present).
FTDNA shows the lone C1e sample coming from Iceland.
C1f is apparently found among Native Americans, Italians and Tajik-speakers from Tajikistan, taking the geographies of samples from both the FTDNA and YFull trees into account. However, according to the Wikipedia article of mtDNA haplogroup C, there is an ancient sample from NW Russia dating to the Mesolithic.
C1g has been found in an ancient sample from Karelia dating to the Mesolithic according to The genetic history of Ice Age Europe (Qiaomei Fu et al. 2016). However I saw on the YFull tree that an ancient C1g dating to 900 ybp has been found in Bolivia, from Ancient mitochondrial DNA provides high-resolution time scale of the peopling of the Americas (Llamas, Haak, et al. 2016). The important thing to note about the age of the Bolivian sample is that it dates to before the European colonization of the New World, so a later migration can be excluded.
The interesting question then is whether or not C1b, C1c, and C1d, the only C1 subclades found exclusively in the Americas, may all in fact descend from a MRCA who lived after the Northeast Asian C1 MRCA but who did not acquire any stable mtDNA mutations.
If this was the case, it is possible that only one woman / her closer relatives introduced C1b-d (before those individual mutations occurred) to the New World but that the mtDNA genetic evidence either does not exist (i.e. no mutations in that ancestral line), has been obscured (i.e. mutations in unstable region have been subsequently overwritten), or will eventually be established by some potential future correction to the mtDNA phylogenetic tree of C1.
It is important to keep in mind that while the Y-chromosome can accumulate a SNP every hundred or so years that will not diminish reproductive success, mtDNA mutations usually result in the death of the embryo. Because of this, thousands of years can go by before a stable SNP is acquired in a particular mtDNA line.
If I’m reading the Llamas, Haak, et al. (2016) paper correctly, then dendrogram Fig 3 Dated Bayesian mitogenomic tree and reconstruction of past effective female population size appears to show C1b and C1d descending from a more recent common ancestor than they have with C1c. Neither YFull nor FTDNA trees show any SNPs uniting these branches, as I summarized in the preceding paragraph, so I’m wondering on what basis the dendrogram computed C1b and C1d to form a clade.
This paper writes that sometime between 24.9 to 18.4 ka, cold conditions forced those living west of the Bering Land Bridge further south, isolating those living east of the Kamchatka and Chukotka Peninsulas in eastern Beringia.
C1g may have already fully formed before the separation of the two populations if we consider it improbable to migrate to Mesolithic Karelia from eastern Beringia. In that case, after the LGM, one branch ended up on the Siberian side, later migrating to Karelia, while the other ended up isolated in eastern Beringia, later migrating to Bolivia after conditions made it possible to cross the Cordilleran and Laurentide ice sheets.
It says above, “FTDNA shows the lone C1e sample coming from Iceland.”
Can you say more about your thoughts on the claims it may be traced to North America?
Thank you.