I am a volunteer in the J-M241 and J2 Haplogroup Resarch Projects (J-Z2453 focus) on FTDNA.
In the capacity of a volunteer researcher I have reconstructed the following patrilineal trees on the basis of comparing rare shared STR markers:
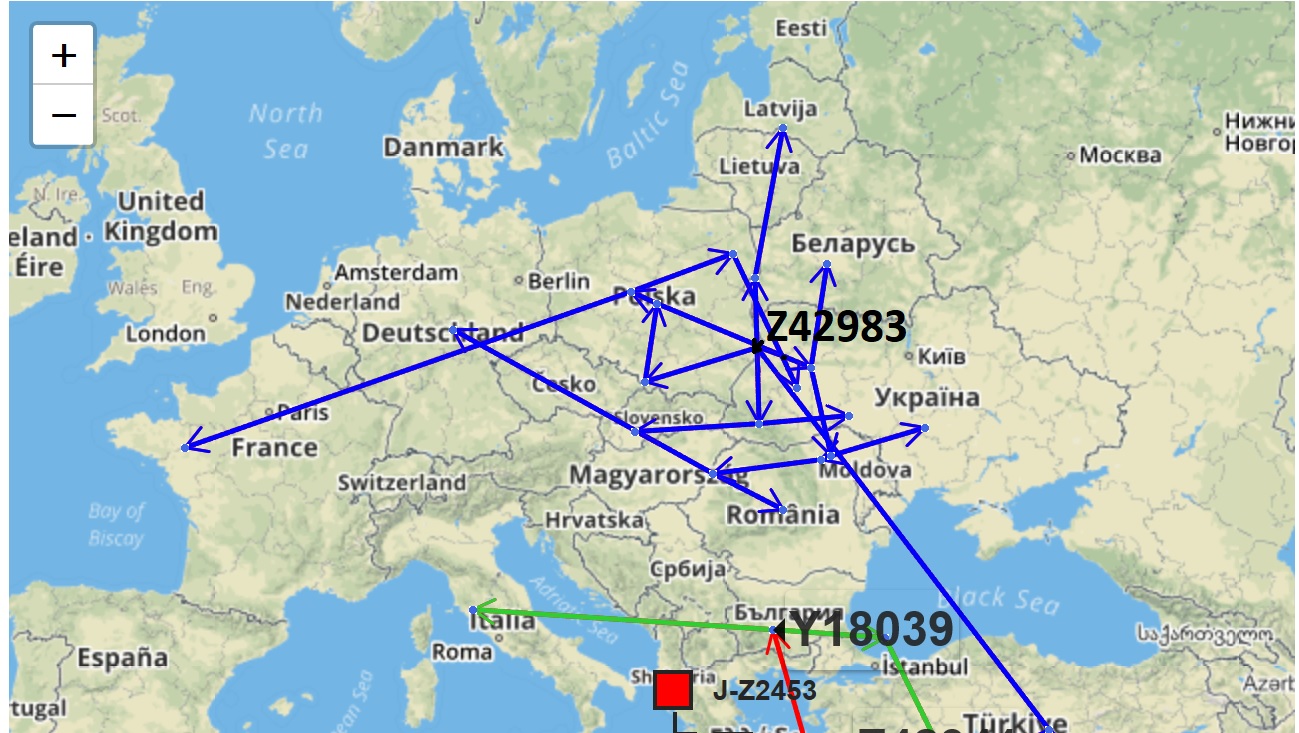
Ashkenazi lineage J-Z42983
These trees are useful for genealogical research because:
- Ignores random noise - Members of your branch in an STR tree are much more likely to share a more recent common ancestor than a group formed purely on the basis of total genetic distance because total genetic distance includes noisy markers. What is noise for one group might not be for another. Analysis reveals this.
- Result can calibrate PhyloGeographer's computed migration path for your lineage.
- What was once a jumbled mess of "Europe" for an entire haplogroup became dissected into regions.
- A more meaningful migration path can now be theorized (below)
I offer this Y-STR analysis as a service to those of other haplogroups where I am not a volunteer.
You receive:
- Tree diagram showing presumed ancestral STRs and mutations that define branching points
- Your group's theoretical migration computed on PhyloGeographer
- Report containing a timeline of the possible cultural identities of your ancient ancestors based on where the computed path leads
What I require:
- Y-STRs of your group members in tabular form
- Predicted haplogroup of your group members
- (optional - for ancestral STRs) Y-STRs of men from sibling haplogroups
- (optional - to compute migration) location of paternal ancestor of each sample
Fees:
- Tree - $10/12/15 per sample (37/67/111 STRs - $75 minimum)
- Ancestral STR derivation - $25 (optional, increases accuracy)
- Migration - $10 per sample ($50 minimum)
Note:
- Ancestral STRs derived only for those markers that my analysis reveals as useful for differentiating members of your haplogroup
- Migrations currently not computed for New World. In a few months after making updates to PhyloGeographer the New World will be possible.
my gmail is hunterprovyn
(+1)434-422-2547
