Here is a brief announcement of the splitting of J1-FGC11, its significance for haplogroup research, and a short explanation of the relative frequency and diversity of this haplogroup we can learn from the Y Frequency and Diversity Heatmaps.
My goal is to explain the tools that can be used to analyze the frequency and diversity of a subclade and leave it to experts in J1 to theorize in earnest about the origin of J1-FGC11, in concert with those who have historical and archaeological expertise.
Yesterday YFull's live tree showed a new split in J1-FGC11, which itself has TMRCA estimated at 1800 BCE.
This subclade has 3205 geolocated samples on the YFull YTree and is found chiefly in the Middle East and North Africa. It accounts for 43.2% of all Saudi Arabia and 37.7% of all Yemen-marked samples on the tree.
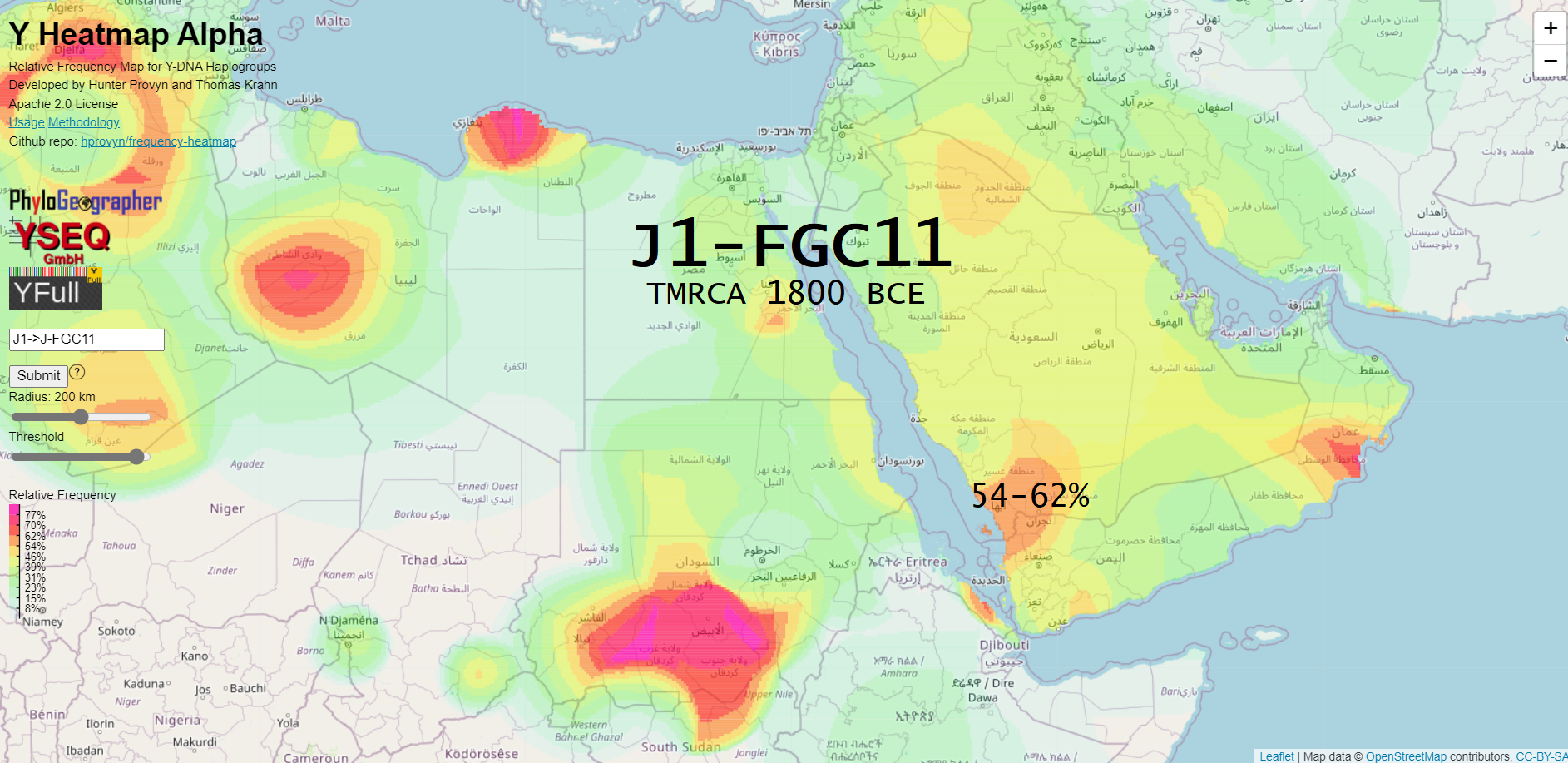
As a fun side note, notice the low percentage on the island of Socotra. This island is mostly inhabited by my distant cousins in J2b-M205. I, being J2b-L283, am more closely related to 80% of the inhabitants of Socotra than they are to most of their neighbors from the Arabian peninsula.
Until now, all the J1-FGC11 men had been positive for two SNPs, FGC11 and Y3014/FGC7638.
Now, after 3205 samples, a man tracing his male line to Dhamar, Yemen has been found to be positive for FGC11 and negative for Y3014/FGC7638.
I wanted to share a few observations regarding the geographic distribution of samples that J1 researchers might take into account in theorizing the origin of J1-FGC11 and related lineages.
Prolific Descendant J1-FGC1707 Has a Much Different Distribution
About 2/3 of all J1-FGC11 on YFull is positive for child J1-FGC1707 with a TMRCA 2500 years ago.
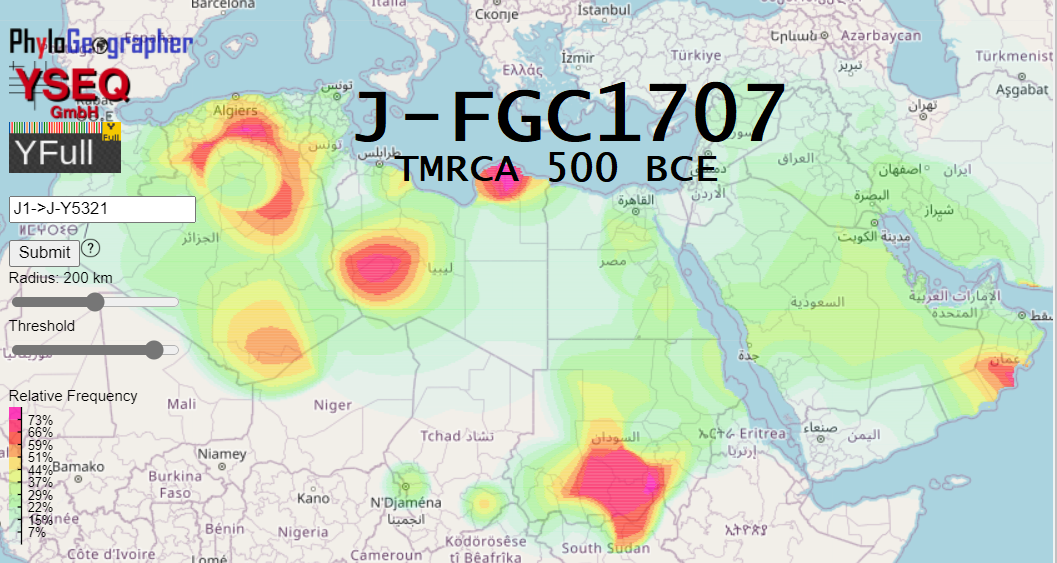
By comparing this frequency map against ancestor J1-FGC11, we see that every local maximum outside of the Arabian peninsula, but also including the one in Oman, is attributed to descendants of the single much younger (Iron Age) descendant, J1-FGC1707.
Notably, the one significant hotspot of J1-FGC11 that is absent in J1-FGC1707 is western Yemen, the Jizan, Asir and Najran regions of Saudi Arabia bordering it, and southern Eritrea.
If we count the J1-FGC11 who are not J1-FGC1707, we get a significantly different ratio between Saudi and Yemen origin men:
660 / 4036 = 16.4% of men tracing to Saudi Arabia are J1-FGC11(xFGC1707)
186 / 546 = 34.1% of men tracing to Yemen are J1-FGC11(xFGC1707)
I plan to develop tools to automatically tabulate these kind of country and region-level statistics from the samples on the YFull tree.
Once these tables are complete, researchers may refer to such statistics computed with greater regional specificity.
After all, the question of the geographic origin of J1-FGC11 is not simply Saudi Arabia vs Yemen or other modern country - these boundaries are more recent political constructs.
The relative frequency hotspot of J1-FGC11, excluding younger J1-FGC1707, is the geography around western Yemen and Jizan, Asir and Najran provinces of Saudi Arabia.
What does the Diversity Map show?
The above example is a great case study for how a relative frequency map of a haplogroup can show areas with greater frequency of a haplogroup that happen to have formed much later, by founder effects of individual men who migrated there from a different, deeper geographic origin.
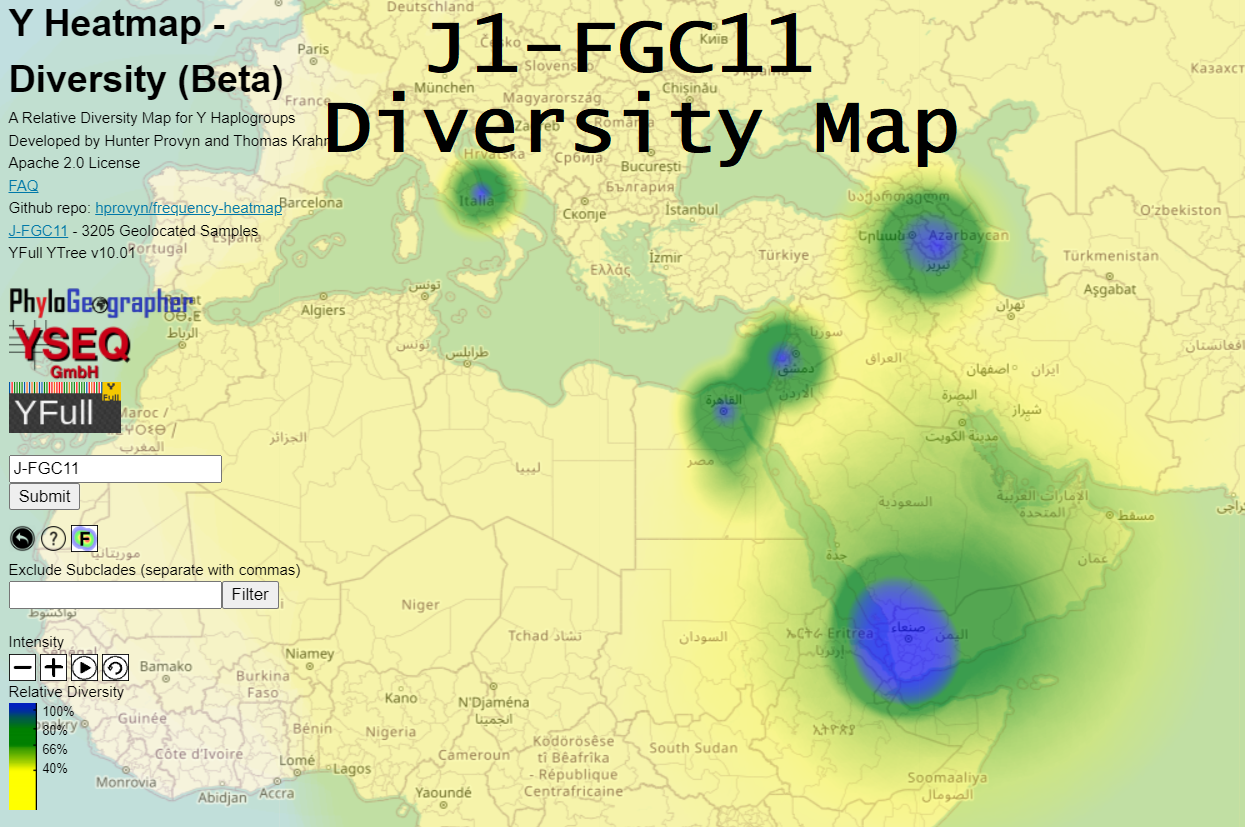
The diversity map for J1-FGC11 (above) indicates very high diversity around a large area of western Yemen and south-western provinces of Saudi Arabia.
It makes sense to adjust the intensity by clicking + or - until you see regional differentiation (i.e. the whole area is not blue).
Note that this diversity map was computed using v10.02, before J1-FGC11 was split. Now the diversity around Yemen will increase even more.
The way diversity is calculated, the single Yemenite sample from Dhamar will count as much as all the other samples downstream of new sibling J1-Y3014 combined.
The further downstream samples are, the less impact they have on the overall diversity. This is why diversity maps are immune from founder effects, which occur further down in the tree from the ancestor.
Conclusion of this short analysis of J1-FGC11
In this very cursory overview I observed that a major local maximum of relative frequency (especially after excluding the major founder effect of J1-FGC1707) aligns with the major local maximum of diversity. This could be indicative of an origin of J1-FGC11 in western Yemen, Jizan, Asir and Najran provinces of Saudi Arabia or an ancient co-migration to this area.
To be sure, more in-depth analysis of each particular branch's distribution, and taking ancient samples into account is warranted.
I think it is also necessary for anyone looking to theorize on the geographic origin of J1-FGC11 to also examine the geographic distribution of its siblings. The siblings are found in the Middle East but also throughout Eurasia. An apple usually doesn't fall too far from the tree.
Further Testing in Western Yemen and in Eritrea, Ethiopia, Djibouti, and Sudan Would Advance the Research Further
I am told by the researcher whose sample from Dhamar split J1-FGC11 that they are only scratching the surface in testing some of the villages in this region of Yemen. I count 21 samples from Dhamar on the last YTree v10.02.
I would also recommend to do more testing in the very undersampled regions on the other side of the Red Sea. Even if these areas on the other side of the Red Sea don't end up contributing much to understanding the deeper origin of J1-FGC11 or J1, it may increase our understanding of some of the haplogroups of A-M32 and E.
Next Steps for this J1-FGC11* Sample
The next step to advance the research into the origin of this parallel line of J1-FGC11 is to find a hopefully distant relative of his that will share some SNPs with him below the J1-FGC11 branching point.
I will use STR Match Finder to recommend possible candidates for testing. It provides a clear visual indication of shared rare alleles that might be indicative of inheritance through common descent.
With more precise geographic origin input data, we can get more precise, possibly even tribal-level origin maps
It is important to understand the limitations of the maps I have presented.
These were computed based on samples whose origins I could observe from the public YFull tree, that is, at the level of resolution of country-regional code.
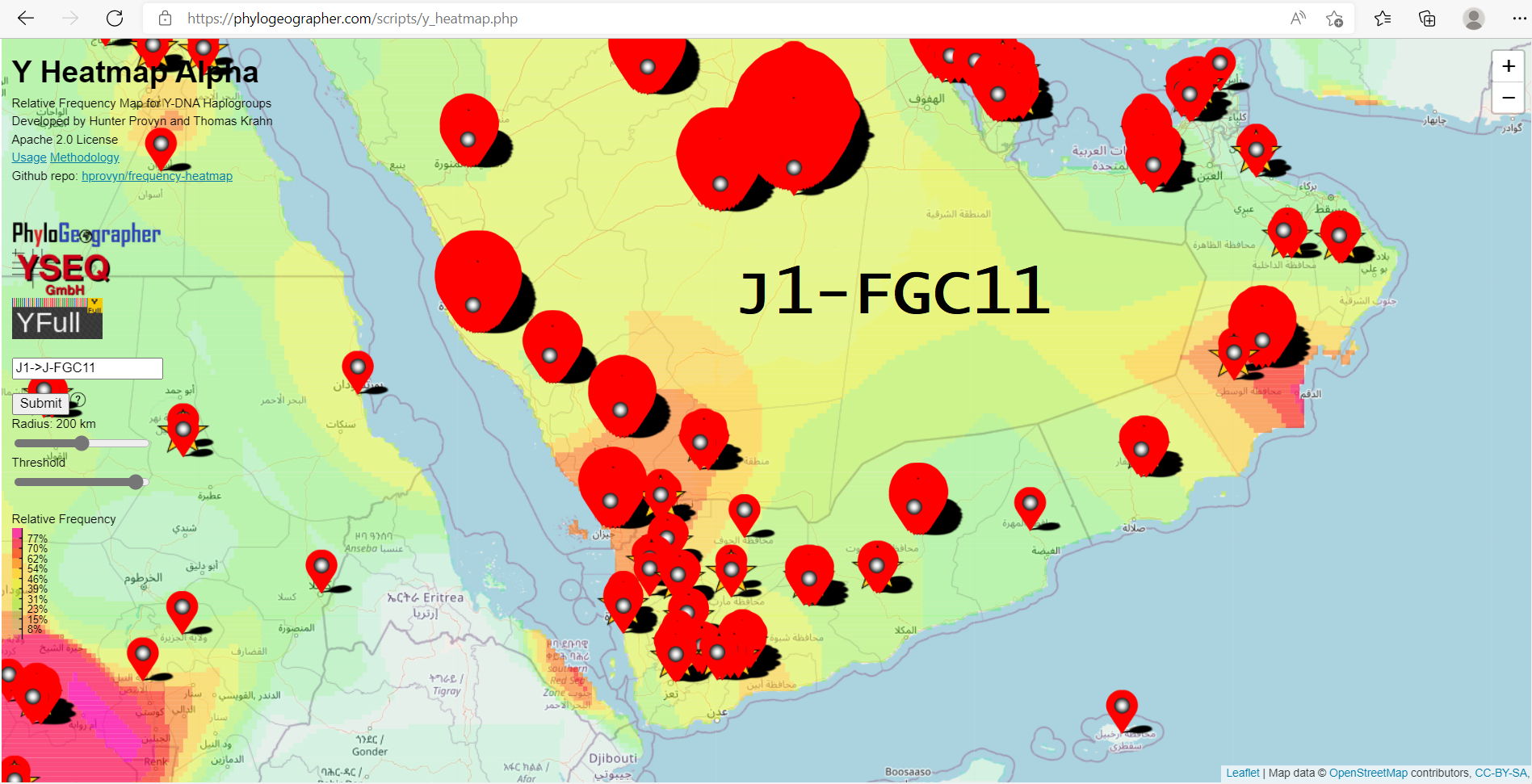
As long as the regions are relatively smaller, like in western Yemen, this is sufficient to compute a gradient that is accurate on the regional level.
However, the distribution in larger regions like Ash Sharqiyah, Saudi Arabia suffers from having its samples being represented by a single location. Sometimes there is no best single position to represent a huge region.
If I can get precise latitude longitudes of the male and female line ancestors of YTree and MTree samples it will facilitate a much more accurate heatmap.
Please do not contact me on an individual basis to supply your YFull coordinates. Any kind of possible collection I can do must be done in a coordinated, scalable manner.
Note About PhyloGeographer Theoretical Computed Paths For Arabian Haplogroups
Please keep in mind that theoretical computed paths from PhyloGeographer are theoretical and do not, on their own, constitute proof of origin or migration path.
It is important to understand that the algorithm I wrote, several years ago, does not take sample rate into account.
Due to the extremely high sample rate of Saudi Arabia compared to most of its neighbors, this means that haplogroups with any presence in Saudi Arabia are likely to be computed as originating there because of the high sample rate and not because they really originated there.
I have other development priorities at this time, including M Heatmap, but if anyone would like to sponsor me to continue to make improvements to PhyloGeographer, they may send me a donation.
Y Heatmap and M Heatmap are supported by YSEQ but PhyloGeographer is my own independent work that has not been financially supported except through modest monthly contributions from my Patreon subscribers.
Subscribers to my PhyloGeographer on Patreon services get Y and M Heatmaps and a link showing their male line ancestor added to the end of their theoretical Y-DNA haplogroup migration path from PhyloGeographer. They also get sneak previews of the new features I develop.
By the way, YSEQ is now offering a higher coverage WGS test for a lower price than before ($359 for 15x, $399 for 30x, $699 for 50x coverage).
Hire Me To Advance Your Own Research Goals
I do Y-DNA consulting to advance your research goals for $50 an hour. Contact me by email to set up a call: hunter provyn at gmail dot com