Use Case
You have a sample that tested STRs at YSEQ, FTDNA or elsewhere.
Later this same sample did a WGS test and you have access to his STRs extracted by YFull.
Now you would like to automatically merge his STR alleles from both tests into one single entry (i.e. one row of a tabular input data source) to use to find matches in STR Match Finder.
How to use it
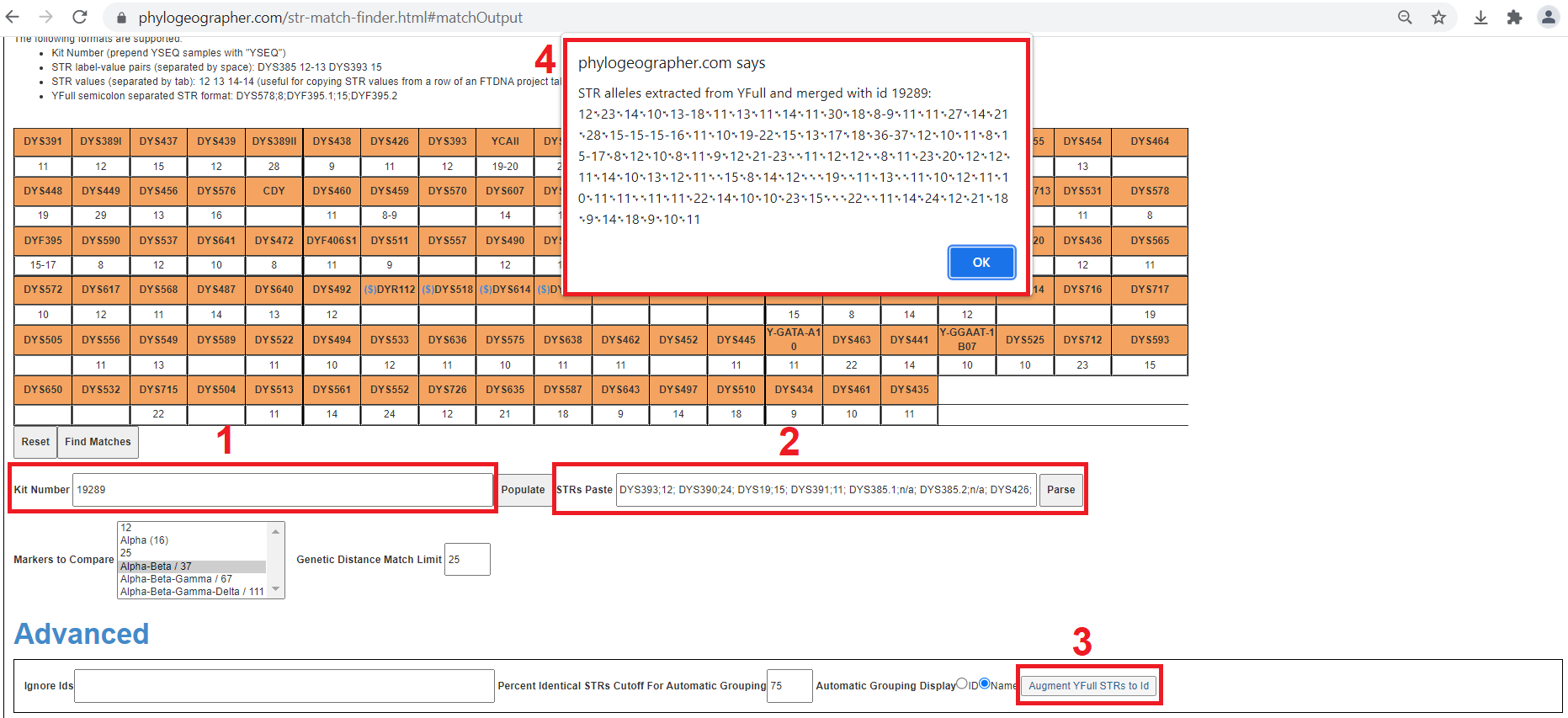
- Enter the id of the sample whose pre-existing STRs (from YSEQ or imported from Tabular Data source) you would like to merge with in the Query->"Kit Number" field. (Optional) Click Populate to ensure you actually have this guy's STRs imported.
- Paste the YFull-extracted STRs (just select "111" option from YFull) into the Query->"STRs Paste" field and click "Parse". This overwrites his STRs in the grid view with those extracted by YFull, but does not merge as desired.
- Click the button in Query->Advanced->"Augment YFull STRs to Id"
- A popup informs you the id (optional) that the YFull-extracted STRs were merged with and provides you with tab-separated alleles you can copy-paste directly into your Tabular Data source.
- If the sample you merged with was a YSEQ sample rather than one you had imported from a Tabular Data source, use this template to create a Tabular Data source
- Note that this process does not merge the STRs currently stored in the webpage. You must reload the page and reimport the updated Tabular Data source to have the merged STRs associated with the id in the system.
I have done the big y [updated to y 700 I think. I have genealogy data in my family tree to about 1769 and dna research of the y side and mitochondrial side back to 1570 and some info to 1500 what would you recommend would be a way to enhance this tree. My kit number is 211269. by40713 and h1j.
Sorry for the late reply, I’ve emailed you just now regarding your query.