With the latest update to version 3 of the PhyloGeographer algorithm, migration paths are now computed to the New World for selected haplogroups within C and Q (C-1373, Q-Z780, Q-M3, Q-F746).
Some paths will now cross the International Date Line which lies in the middle of the Pacific Ocean. This is determined by the algorithm which now uses great-circle distance and only averages latitudes and longitudes after first applying an offset to avoid undesirable behavior caused by -180° being equivalent to 180°.
For example, without applying any offset, 178° and -176° would average to 1°. This is clearly not a good estimator for the center of these two points. By application of an offset the center is calculated at -179°, which is just 3° distant from each.
How did Q turn out?
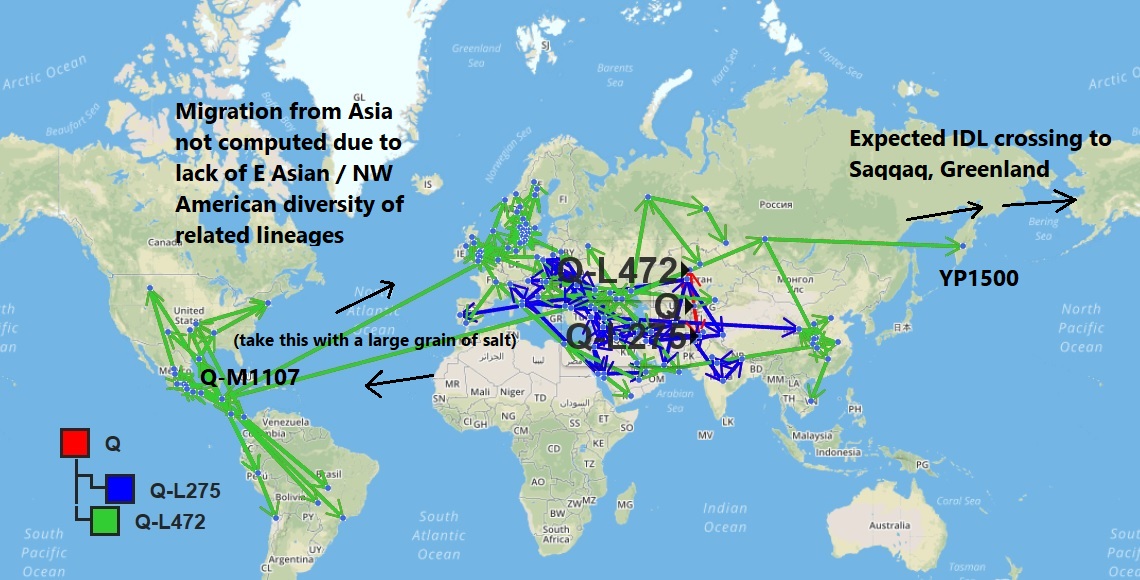
A migration to the New World via Asia was computed for YP1500 sample Saqqaq, but not for Q-M1107.

The algorithm computed the path of YP1500 from Kamchatka eastward to Greenland as expected. However, the shortest great-circle path between YFull samples of M1107 and L330 crosses the Atlantic rather than the Pacific.
This should change if/when more L330 is found in North East Asia, some M1107>M930>L804 is found in Eurasia (presently only Scandinavia/England) and more early M930 diversity is found in NW America.
Each of these lineages is diverse enough to warrant its own article. Once we have more data and the algorithm is more robust I will revisit. In the meantime any comments are welcome on this thread.
M1107 being computed to Central America could be accurate or the result of the extinction of other NW American M1107 lineages.
As always take any computed path with a grain of salt, especially these which are computed on samples which have diverged thousands of years ago and crossed great distances.
The PhyloGeographer algorithm is a work in progress and now open source. Suggestions for improvements are welcome.
Absolutely monumental work if we are to extrapolate that ancient Native Americans traversed the Florida Current which would explain why mtDNA X is also found in the the mound builder culture of Newgrange, from 9k yo bog bodies of FL? The tremendous complexity of these ancient people’s is remarkable whichever way they spread the globe with a unifying culture of mound and tell building, black earths as well as a mythology that permeates every major civilization the world wide and also extremely ancient! If haplogroup R-V88 turns up in an ancient sample from South America and considering that R is thought to have traveled along side Q during deep antiquity from it’s split from P, we can be fairly certain of the possibility that they could have traversed the Brazil Current to Cameroon according to this algorithm? Completely fascinating to even say the least! Thank you!