mt Clade Finder is an mtDNA analytical tool developed by Hunter Provyn and Thomas Krahn of YSEQ that takes a FASTA file as input.
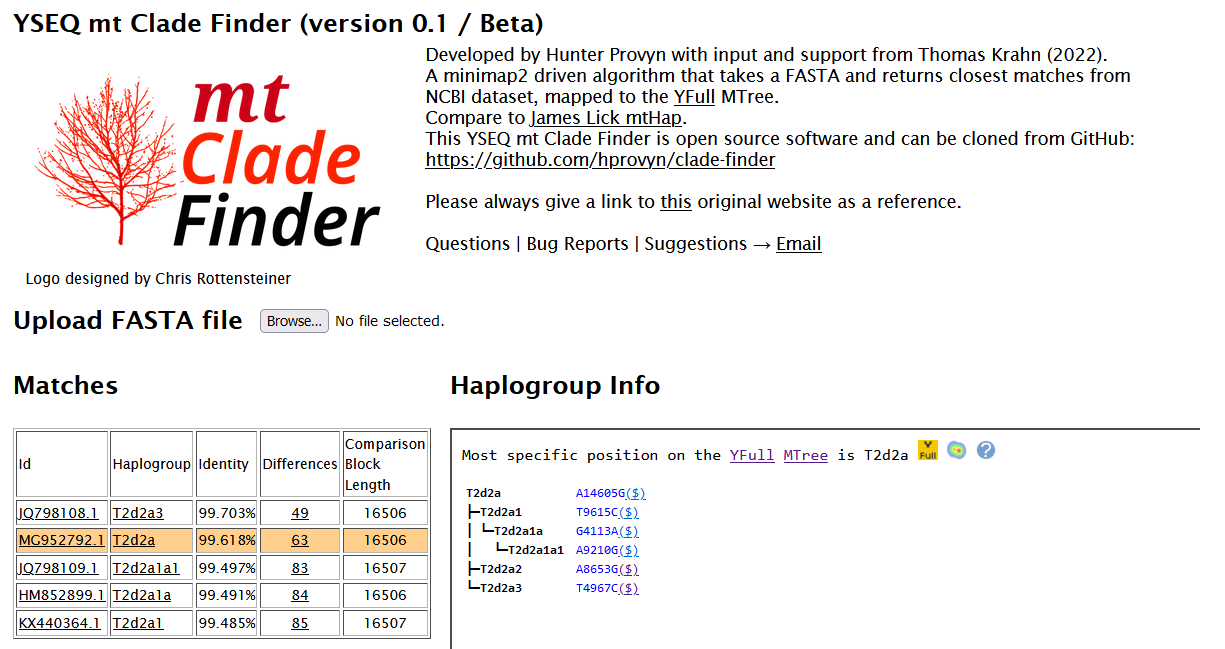
The Matches table provides information and links to your closest matches from samples within the NCBI database that have been analyzed by the YFull MTree.
Haplogroup Info section depicts a simplified tree view of the relevant branch of the YFull MTree. Each branch lists the mtDNA SNPs and their corresponding panel at YSEQ. At the top are clickable icons that link to that branch on the MTree and its relative frequency across the world as computed by mt Heatmap.
By clicking on the Differences number for one of your matches, a more detailed Differences to rCRS loads below.
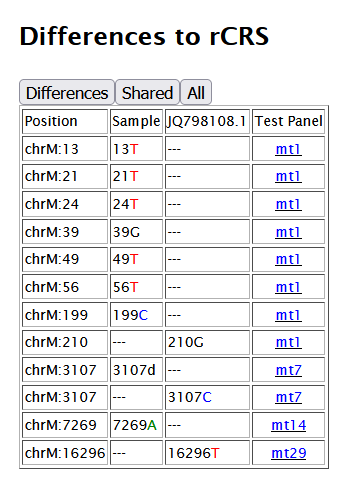
Links to the corresponding mtPanel test are provided so one may confirm whether or not they share a mutation with their NCBI match which may or may not be indicated on the YFull MTree.
There are buttons to filter the view between the differences unique to each sample vs shared by both.
Benefits of YFull Analysis
This is a tool for analyzing a mtDNA FASTA but is not intended as a replacement for having your mtDNA FASTA analyzed on YFull.
Though we have not found any errors in our testing with complete mitochondrial sequences, this software is automated, backed by minimap2, and not guaranteed to always produce the correct result. The tool might accept shorter fasta files, but there's no warranty that this will deliver any useful interpretation.
As this tool does not store your data, it does not contribute to the research by adding a new branch to the public research tree in the way that YFull analysis does.
The accuracy of this tool's haplogroup prediction becomes more powerful and the linked mt Heatmap relative frequency maps become more accurate the more samples are added to the YFull MTree.
Is the mt clade-finder going to be receiving an update so that tests like 23andme and FamilyTreeDNA can be uploaded, similar to what the Y clade-finder allows, instead of only allowing fasta files?
This is not currently planned.
mtComplete is one product that will provide a FASTA of the complete mtDNA – https://www.yseq.net/product_info.php?cPath=28&products_id=38291